Publication
-
Single-Cell RNA-Seq Reveals Conserved Cellular Communication Mechanisms Governing Ocular Lineage Specification from Human iPS Cells
Laura Howard, Yuki Ishikawa, Rei Kamuro, Tomohiko Katayama, Kiranjit K. Bains, Matthew J. Hill, Derek J. Blake, Sung-Joon Park, Ryuhei Hayashi, Andrew J. Quantock, Kohji Nishida
Cells 15:104 (2026) -
Machine learning-based predictive modeling maximizes the efficacy of mTOR/p53 co-targeting therapy against AML
Jingmei Li, Emi Sugimoto, Keita Yamamoto, Yutong Dai, Wenyu Zhang, Yu-Hsuan Chang, Jakushin Nakahara, Tomohiro Yabushita, Toshio Kitamura, Sung-Joon Park, Kenta Nakai, Susumu Goyama
Cancer Science 116:3040-3051 (2025) [PUBMED] -
RNA secondary structure prediction by conducting multi-class classifications
Jiyuan Yang, Kengo Sato, Martin Loza, Sung-Joon Park & Kenta Nakai
Computational and Structural Biotechnology Journal 27:1449-1459 (2025) [PUBMED] -
STAIG: Spatial transcriptomics analysis via image-aided graph contrastive learning for domain exploration and alignment-free integration
Yitao Yang, Yang Cui, Xin Zeng, Yubo Zhang, Martin Loza, Sung-Joon Park & Kenta Nakai
Nature Communications 16:1067 (2025) [PUBMED] [Press Release] [EurekAlert]
STAIGは、空間トランスクリプトーム解析において、画像情報を活用して高精度な空間ドメインの同定を可能にする新しい深層学習モデルです。本モデルでは、各スポットにおける画像から自己教師あ>り学習手法により特徴ベクトルを抽出し、これを遺伝子発現データと統合することで、空間的コンテキストをより的確に捉えることが可能となります。このアプローチにより、従来手法では見落とされがちであった微細な空間構造の検出や、組織内の機能的領域の同定が向上します。 -
In-silico analysis revealed Marco (SR-A6) and Abca1/2 as potential regulators of lipid metabolism in M1 macrophage hysteresis
Yubo Zhang, Wenbo Yang, Yutaro Kumagai, Lopez Martin, Yitao Yang, Sung-Joon Park & Kenta Nakai
International Journal of Molecular Sciences 26:111 (2025) [PUBMED] Single-cell transcriptomics reveals the molecular basis of human iPS cell differentiation into ectodermal ocular lineages
Laura Howard, Yuki Ishikawa, Tomohiko Katayama, Sung-Joon Park, Matthew J. Hill, Derek J. Blake, Kohji Nishida, Ryuhei Hayashi & Andrew J. Quantock
Communications Biology 7:1495 (2024) [PUBMED]Computational analysis of the functional impact of MHC-II-expressing triple-negative breast cancer
Yang Cui, Weihang Zhang, Xin Zeng, Yitao Yang, Sung-Joon Park and Kenta Nakai
Frontiers in Immunology 15:1497251 (2024) [PUBMED]
腫瘍微小環境(TME)は腫瘍の進行と免疫調節に重要な役割を果たします。MHCクラスII分子はTME内で免疫監視に必要不可欠であり、通常は抗原提示細胞のみ発現しますが、腫瘍細胞にも発現することがあります。本研究では、乳がんのバルク、単一細胞、空間的遺伝子発現データを解析し、MHC-IIを発現する腫瘍細胞が免疫細胞と相互作用し、良好な予後群に関連する免疫浸潤部位に存在する傾向を見い出しました。この知見は、MHC-II発現腫瘍細胞が免疫環境の調整において重要であり、患者の生存予後に影響を与える可能性があることを示唆しています。TF-EPI: an interpretable enhancer-promoter interaction detection method based on transformer
Bowen Liu, Weihang Zhang, Xin Zeng, Martin Loza, Sung-Joon Park and Kenta Nakai
Frontiers in Genetics 15:1444459 (2024) [PUBMED]
この研究では、DNA配列を用いてエンハンサーとプロモーターの相互作用(EPI)を検出するための新しい深層学習モデルを開発しています。特に、転移学習を導入することで、細胞タイプ特異的な遺伝子調節に関連するEPIの検出精度が向上しました。この進展により、エンハンサーとプロモーターからの転写因子の作用をより深く分析できるようになり、遺伝子調節機構の多様性と共通性の理解が進むことが期待されます。Integrative analysis of cancer multimodality data identifying COPS5 as a novel biomarker of diffuse large B-cell lymphoma
Yutong Dai, Jingmei Li, Keita Yamamoto, Susumu Goyama, Martin Loza, Sung-Joon Park and Kenta Nakai
Frontiers in Genetics 15:1407765 (2024) [PUBMED]A computational approach for deciphering the interactions between proximal and distal gene regulators in GC B-cell response
Sung-Joon Park* and Kenta Nakai
NAR Genomics and Bioinformatics 6:lqae050 (2024) [PUBMED]
この研究では、遺伝子発現の調節に関与する転写メディエーター複合体の機能を理解するために、プロモータ近接および遠位調節因子の相互作用をモデル化する計算手法を開発しました。回帰モデルを用いて重要な調節因子を特定し、グラフ埋め込み手法で共同調節された遺伝子を検出することで、ヒト胚中心B細胞の遺伝子調節メカニズムを詳細に調査しました。結果、プロモータ近接調節因子が主に遺伝子発現を制御する一方で、遠位調節因子は転写を微調整することが明らかになりました。また、共発現するコファクターや近接・遠位転写因子などのモジュール型調節因子が存在することが分かり、一部のモジュールはリンパ腫において異常な発現パターンを示しました。この結果は、転写因子と構造因子の相互作用の調節不全がクロマチン再構成の失敗に関連し、悪性腫瘍のリスクを高める可能性があることを示唆しています。私たちの計算手法は、空間的に相互作用する転写調節因子のコードを解読するのに役立つと考えられます。HyGAnno: Hybrid graph neural network-based cell type annotation for single-cell ATAC sequencing data
Weihang Zhang, Yang Cui, Bowen Liu, Martin Loza, Sung-Joon Park and Kenta Nakai
Briefings in Bioinformatics 25:bbae152 (2024) [PUBMED]
scATAC-seqデータに対して細胞タイプを注釈するために、scRNA-seqデータにおける細胞タイプ情報を活用する半教師あり学習手法を開発しました。この手法は、遺伝子発現だけでなく、全ゲノムのアクセシビリティ情報も取り入れることで、細胞注釈の信頼性を向上させています。これにより、scATAC-seqデータの細胞タイプ注釈がより正確に行えるようになります。Genome-wide ATAC-see screening identifies TFDP1 as a modulator of global chromatin accessibility
Satoko Ishii, Taishi Kakizuka, Sung-Joon Park, Ayako Tagawa, Chiaki Sanbo, Hideyuki Tanabe, Yasuyuki Ohkawa, Mahito Nakanishi, Kenta Nakai and Yusuke Miyanari
Nature Genetics 56:473-482 (2024) [PUBMED] [Press Release]
本研究では、全ゲノムCRISPRスクリーニングとATAC-seeプロトコルを組み合わせ、細胞レベルでクロマチン状態に影響を与える遺伝子を特定しました。膨大なシークエンシングデータ解析により、既知の調節因子に加え、TFDP1やHNRNPUなどの新たなタンパク質を発見し、それぞれが異なる影響を持つことを確認しました。特に、TFDP1はヒストン転写調節を通じてクロマチン状態を間接的に調整することが明らかとなり、細胞工学やゲノム編集への応用が期待されます。Multi-omics computational analysis unveils the involvement of AP-1 and CTCF in hysteresis of chromatin states during macrophage polarization
Yubo Zhang, Wenbo Yang, Yutaro Kumagai, Martin Loza, Weihang Zhang, Sung-Joon Park and Kenta Nakai
Frontiers in Immunology 14:1304778 (2023) [PUBMED]Signaling Pathway of Taurine-Induced Upregulation of TXNIP
Hideo Satsu, Yusuke Gondo, Hana Shimanaka, Masato Imae, Shigeru Murakami, Kenji Watari, Shunichi Wakabayashi, Sung-Joon Park, Kenta Nakai & Makoto Shimizu
Metabolites 12:636 (2022) [PUBMED]Generation of 3D lacrimal gland organoids from human pluripotent stem cells
Ryuhei Hayashi, Toru Okubo, Yuji Kudo, Yuki Ishikawa, Tsutomu Imaizumi, Kenji Suzuki, Shun Shibata, Tomohiko Katayama, Sung-Joon Park, Robert D. Young, Andrew J. Quantock & Kohji Nishida
Nature 605:126-131 (2022) [PUBMED] [Press Release]Discovery of widespread transcription initiation at microsatellites predictable by sequence-based deep neural network
Mathys Grapotte, Manu Saraswat, Chloé Bessière, Christophe Menichelli, Jordan A. Ramilowski, Jessica Severin, Yoshihide Hayashizaki, Masayoshi Itoh, Michihira Tagami, Mitsuyoshi Murata, Miki Kojima-Ishiyama, Shohei Noma, Shuhei Noguchi, Takeya Kasukawa, Akira Hasegawa, Harukazu Suzuki, Hiromi Nishiyori-Sueki, Martin C. Frith, FANTOM consortium (incl. Sung-Joon Park, Kenta Nakai), Clément Chatelain, Piero Carninci, Michiel J. L. de Hoon, Wyeth W. Wasserman, Laurent Bréhélin & Charles-Henri Lecellier
Nature Communications 12:3297 (2021) [PUBMED]Characterizing promoter and enhancer sequences by a deep learning method
Xin Zeng, Sung-Joon Park and Kenta Nakai
Frontiers in Genetics 12:681259 (2021) [PUBMED]OpenContami: A web-based application for detecting microbial contaminants in next-generation sequencing data
Sung-Joon Park* and Kenta Nakai
Bioinformatics 37(18):3021-3022 (2021) [PUBMED]
細胞実験における微生物の汚染は研究結果に甚大な影響を与えるため、その迅速かつ定量的評価は極めて重要です。この問題を解決するために「OpenContami」というオンラインアプリケーションを開発しました。このアプリは、次世代シーケンシング(NGS)データを使って、外来微生物種を簡単に検出できるツールであり、大規模公共データセットからの分析結果やユーザーがアップロードしたデータを含むデータベースが組み込まれています。OpenContamiを使うことで、研究者はNGSデータに含まれる微生物の影響をより深く理解でき、実験の正確性と信頼性を向上させることができます。A semi-supervised deep learning approach for predicting the functional effects of genomic non-coding variations
Hao Jia, Sung-Joon Park and Kenta Nakai
BMC Bioinformatics 22(Suppl 6):128 (2021) [PUBMED]
この研究では、非コーディング領域の変異が遺伝子発現や病気に与える影響を予測するため、半教師あり深層学習モデルを提案しています。実験データと注釈なしデータを組み合わせて学習させ、従来のツールより優れた予測性能を実現しました。また、非コーディング変異が細胞特異的であること、DNAのアクセシビリティが変異の影響に重要であることが明らかになりました。この方法は、限られたデータを最大限活用し、病気に関連する変異を特定する新しい手法を提供します。Existence and possible roles of independent non-CpG methylation in the mammalian brain
Jong-Hun Lee, Yutaka Saito, Sung-Joon Park, Kenta Nakai
DNA Research 27:dsaa020 (2020) [DNA Res] [PUBMED]RNA-sequencing reveals positional memory of multipotent mesenchymal stromal cells from oral and maxillofacial tissue transcriptomes
Satoru Onizuka, Yasuharu Yamazaki, Sung-Joon Park, Takayuki Sugimoto, Yumiko Sone, Sebastian Sjöqvist, Michihiko Usui, Akira Takeda, Kenta Nakai, Keisuke Nakashima, Takanori Iwata
BMC Genomics 21, 417 (2020) [BMC] [PUBMED]HHEX promotes myeloid transformation in cooperation with mutant ASXL1
Reina Takeda, Shuhei Asada, Sung-Joon Park, Akihiko Yokoyama, Hans Becker, Akinori Kanai, Valeria Visconte, Courtney Hershberger, Yasutaka Hayashi, Taishi Yonezawa, Moe Tamura, Tsuyoshi Fukushima, Yosuke Tanaka, Tomofusa Fukuyama, Akiko Matsumoto, Satoshi Yamasaki, Kenta Nakai, Satoshi Yamazaki, Toshiya Inaba, Tatsuhiro Shibata, Daichi Inoue, Hiroaki Honda, Susumu Goyama, Jaroslaw Maciejewski, and Toshio Kitamura
Blood, blood.2019004613 (2020) [Blood] [PUBMED]A systematic sequencing-based approach for microbial contaminant detection and functional inference
Sung-Joon Park, Satoru Onizuka, Masahide Seki, Yutaka Suzuki, Takanori Iwata, and Kenta Nakai
BMC Biology (2019), 17(1):72 [BMC] [PUBMED]
細胞実験における微生物の汚染がデータ分析に与える影響を解析するために、次世代シーケンシング(NGS)データから外来性微生物を特定する計算手法を提案しました。ここでは、1百万の宿主RNA-seqリードあたり1,000から100,000の微生物リードが検出されることを明らかにしました。特に、Cutibacteriumが一般的な汚染因子であることが示され、汚染は主に実験室環境から由来する可能性が高いことがわかりました。また、微生物汚染によって宿主細胞のフェノタイプに重要な変化が生じることも明らかになりました。この研究は、微生物汚染の正確な評価が関連研究の質を高めるために重要であることを強調しています。Broad heterochromatic domains open in gonocyte development prior to de novo DNA methylation
Soichiro Yamanaka, Hidenori Nishihara, Hidehiro Toh, Luis Augusto Eijy Nagai, Kosuke Hashimoto, Sung-Joon Park, Aoi Shibuya, Ana Maria Suzuki, Yujiro Tanaka, Kenta Nakai, Piero Carninci, Hiroyuki Sasaki, and Haruhiko Siomi
Dev. Cell (2019), S1534-5807(19)30629-X [Dev.Cell] [PUBMED]Analyzing the 3D chromatin organization coordinating with gene expression regulation in B-cell lymphoma
Luis A.E. Nagai, Sung-Joon Park and Kenta Nakai
BMC Med Genomics (2019), 11 (Suppl 7):127 [BMC] [PUBMED]Differential landscape of non-CpG methylation in embryonic stem cells and neurons caused by DNMT3s
Jong-Hun Lee, Sung-Joon Park and Kenta Nakai
Scientific Reports (2017), 7(1):11295 [SREP] [PUBMED]
この研究では、胚性幹細胞と神経細胞における非CpG部位(mCpH)のメチル化パターンの特異性を計算的に解析しています。これらのメチル化は、DNAメチルトランスフェラーゼ(DNMT3a、DNMT3b)の細胞特異的な働きによって形成されることが知られています。ここでは、特にESCsでDNMT3bが転写活性の高い遺伝子周辺にメチル化を誘導し、発生に重要な遺伝子を調節していることがわかりました。この研究は、mCpHメチル化が細胞の機能や発展にどのように関与しているかを解明する手がかりを提供します。Multidisciplinary insight into clonal expansion of HTLV-1-infected cells in adult T-cell leukemia via modeling by deterministic finite automata coupled with high-throughput sequencing
Amir Farmanbar, Sanaz Firouzi, Sung-Joon Park, Kenta Nakai, Kaoru Uchimaru and Toshiki Watanabe
BMC Med Genomics (2017), 10(1):4 [BMC] [PUBMED]Transcriptional regulation of a horizontally transferred gene from bacterium to chordate
Yasunori Sasakura, Yosuke Ogura, Nicholas Treen, Rui Yokomori, Sung-Joon Park, Kenta Nakai, Hidetoshi Saiga, Tetsushi Sakuma, Takashi Yamamoto, Shigeki Fujiwara and Keita Yoshida
Proc. R. Soc. B (2016), 283:20161712 [RSPB] [PUBMED]ZBTB16 as a Downstream Target Gene of Osterix Regulates Osteoblastogenesis of Human Multipotent Mesenchymal Stromal Cells
Satoru Onizuka, Takanori Iwata, Sung-Joon Park, Kenta Nakai, Masayuki Yamato, Teruo Okano and Yuichi Izumi
J. Cell. Biochem. (2016), 117:2423-2434 [JCB] [PUBMED]Advances, Practice, and Clinical Perspectives in High-throughput Sequencing
Sung-Joon Park*, Mihoko Saito-Adachi, Yusuke Komiyama and Kenta Nakai
Oral Diseases (2016), 22:353-364 [ODI] [PUBMED]PAX6 isoforms, along with reprogramming factors, differentially regulate the induction of cornea-specific genes
Yuzuru Sasamoto, Ryuhei Hayashi, Sung-Joon Park, Mihoko Saito-Adachi, Yutaka Suzuki, Satoshi Kawasaki, Andrew Quantock, Kenta Nakai, Motokazu Tsujikawa and Kohji Nishida
Scientific Reports (2016), 6:20807 [SREP] [PUBMED]OpenTein: a database of digital whole-slide images of stem cell-derived teratomas
Sung-Joon Park, Yusuke Komiyama, Hirofumi Suemori, Akihiro Umezawa and Kenta Nakai
Nucl. Acids Res. (2016), 44(D1):D1000-D1004 [NAR] [PUBMED] [OpenTein]An integrative approach for efficient analysis of whole genome bisulfite sequencing data
Jong-Hun Lee, Sung-Joon Park and Kenta Nakai
BMC Genomics (2015), 16(Suppl 12):S14 [BMC] [PUBMED]DBTMEE: a database of transcriptome in mouse early embryos
Sung-Joon Park, Katsuhiko Shirahige, Miho Ohsugi and Kenta Nakai
Nucl. Acids Res. (2015), 43(D1):D771-D776 [NAR] [PUBMED] [DBTMEE]Computational promoter modeling identifies the modes of transcriptional regulation in hematopoietic stem cells
Sung-Joon Park, Terumasa Umemoto, Mihoko Saito-Adachi, Yoshiko Shiratsuchi, Masayuki Yamato and Kenta Nakai
PLoS One 9(4):e93853, 2014 [PUBMED]Inferring the choreography of parental genomes during fertilization from ultralarge-scale whole-transcriptome analysis
Sung-Joon Park, Makiko Komata, Fukashi Inoue, Kaori Yamada, Kenta Nakai, Miho Ohsugi and Katsuhiko Shirahige
Genes Dev (2013), 27:2736-2748 [PUBMED] [Press Release] [BRIC] [RESEARCH HIGHLIGHTS]
哺乳類の初期胚発生における分子基盤を理解するため、超大規模なマウス初期胚のトランスクリプトーム解析を行い、超高次元データ解析を実施しました。その結果、母性由来の転写産物や受精直後の転写開始を含む精密な遺伝子プログラムを解読し、重要転写因子の同定に成功しました。本研究は、発生生物学および幹細胞研究における遺伝子転写の体系的なバイオソースを提供した点で高く評価されました。Predicting promoter activities of primary human DNA sequences
Takuma Irie, Sung-Joon Park, Riu Yamashita, Masahide Seki, Tetsushi Yada, Sumio Sugano, Kenta Nakai and Yutaka Suzuki
Nucl. Acids Res. (2011), 39(11):e75 [PUBMED]A regression analysis of gene expression in ES cells reveals two gene classes that are significantly different in epigenetic patterns
Sung-Joon Park* and Kenta Nakai
BMC Bioinformatics (2011), 12 (Suppl 1):S50 [BMC] [PUBMED]Probabilistic Graphical Modeling for Large-scale Combinatorial Regulation of Transcription Factors
Sung-Joon Park, Natsuhiro Ichinose and Tetsushi Yada
Proc. the workshop on Knowledge, Language, and Learning in Bioinformatics (KLLBI), 72-86, 2008Applying a Grid Technology to Protein Structure Predictor ROKKY
Kazutoshi Fujikawa, Wenzhen Jin, Sung-Joon Park, Tadaomi Furuta, Shoji Takada, Hiroshi Arikawa, Susumu Date and Shinji Shimojo
Stud Health Technol Inform. (Proc. HealthGrid 2005), 112:27-36, 2005 [PUBMED]A Study of Fragment-Based Protein Structure Prediction: Biased Fragment Replacement for Searching Low-energy Conformation
Sung-Joon Park
Genome Informatics, 16(2):104-113, 2005 [PUBMED]A Computing System for Protein Structure Prediction with Trial-and-error Process
Hiroshi Arikawa,Shingo Masuda, Tadaomi Furuta,Wenzhen Jin, Sung-Joon Park, Shoji Takada, Kazutoshi Fujikawa and Hideki Sunahara
ACS Transaction, IPSJ, 46(11):407-419, 2005 [CiNii]Two-layered Comparison of Protein Structures by Real-coded GA
Sung-Joon Park, Shoji Takada and Masayuki Yamamura
IPSJ Journal, 46(3):898-910, 2005 [CiNii]GA-based Generic Method for Protein Structure Comparison
Sung-Joon Park and Masayuki Yamamura
Proc. The 2003 IEEE Congress on Evolutionary Computation (CEC 2003), 1528-1535, 2003 [IEEE Xplore]Two-layer Protein Structure Comparison
Sung-Joon Park and Masayuki Yamamura
Proc. The 15th IEEE International Conference on Tools with Artificial Intelligence (ICTAI 2003), 435-440, 2003Asynchronous Real-coded Genetic Algorithm for Simultaneous Protein Structure-based Alignment
Sung-Joon Park and Masayuki Yamamura
Proc. Genetic and Evolutionary Computation Conference (GECCO-2003) Graduate Student Workshop, 276-279, 2003Real-coded Genetic Algorithm to Reveal Biological Significant Sites of Remotely Homologous Proteins
Sung-Joon Park and Masayuki Yamamura
Genetic and Evolutionary Computation Conference (GECCO-2003), 1602-1603, 2003 [Springer]
|
"" | ||

|
| |

|
| |

|
| |

|
| |

|
| |

|
| |

|
| |

|
| |

|
| |

|
| |

|
| |

|
| |

|
| |

|
| |

|
| |

|
| |

|
| |

|
| |

|
| |

|
| |

|
| |

|
| |

|
| |

|
| |

|
| |

|
| |

|
| |

|
| |

|
| |

|
| |

|
| |

|
| |

|
| |

|
| |

|
| |

|
| |

|
| |

|
| |

|
| |

|
| |

|
| |
 |
| |
 |
| |
 |
| |
 |
| |
 |
| |
 |
|
-
Genome-wide scanning of gene expression
Sung-Joon Park and Kenta Nakai
in Encyclopedia of Bioinformatics and Computational Biology (2nd Edition), Elsevier, ISBN 978-0-323-95503-4, 5:186-196, 2025 -
Machine Learning-Based Predictive Modeling Maximizes the Efficacy of mTOR/TP53 Co-Targeting Therapy Against Acute Myeloid Leukemia
Jingmei Li, Emi Sugimoto, Keita Yamamoto, Yutong Dai, Sung-Joon Park, Kenta Nakai, Toshio Kitamura, Susumu Goyama
Blood, 144 (Supplement 1):272, 2024 -
NGSによる細胞製品の有効性及び安全性の評価
朴 聖俊
特集:歯根膜由来細胞シートによる歯周組織再生治療, Precision Medicine (北隆館), 7(1):20-23, 2024 -
タウリンによるTXNIP転写活性亢進機構の解析
薩 秀夫, 権藤 祐輔, 嶋中 花, 井前 正人, 村上 茂, 和多利 研二, 若林 俊一, 朴 聖俊, 中井 謙太, 清水 誠
タウリンリサーチ, 8:11-13, 2022 -
スパイクイン,NGS定量標準化の「量叉」
朴 聖俊
実験医学別冊 「クロマチン解析実践プロトコール」(羊土社), 編集:大川恭行, 宮成悠介, pp 22-24, 2020 -
Homeobox Transcription Factor Hhex Promotes Myeloid Leukemia in Cooperation with Mutant ASXL1
Reina Takeda, Shuhei Asada, Sung-Joon Park, Akihiko Yokoyama, Akinori Kanai, Valeria Visconte, Courtney Hershberger, Yasutaka Hayashi, Taishi Yonezawa, Moe Tamura, Tomofusa Fukuyama, Akiko Matsumoto, Satoshi Yamasaki, Kenta Nakai, Toshiya Inaba, Tatsuhiro Shibata, Daichi Inoue, Hiroaki Honda, Susumu Goyama, Jaroslaw P. Maciejewski, Toshio Kitamura
Blood, 134(Supplement 1):2525, 2019 -
Genome-wide scanning of gene expression
Sung-Joon Park*
in Encyclopedia of Bioinformatics and Computational Biology, Academic Press, 3:452-462, 2019 [ScienceDirect] -
Chapter 7: Computational Inference of Gene Regulation from Whole-Transcriptome Analysis of Early Embryos
Sung-Joon Park and Kenta Nakai
in (I. Ivanov, X. Qian and R. Pal ed.) Emerging Research in the Analysis and Modeling of Gene Regulatory Networks, pp. 241-279, IGI Global, 2016 DOI: 10.4018/978-1-5225-0353-8.ch007, [Sample PDF] -
マウス初期胚を用いた大規模トランスクリプトーム解析
井上 玄志, 朴 聖俊, 古俣 麻希子, 山田 かおり, 岸本 健雄, 中井 謙太, 白髭 克彦, 大杉 美穂
Journal of Mammalian Ova Research 31(2):5111-5111, 2014 [PDF] -
PAX6 isoforms, along with reprogramming factors, differentially regulate the induction of cornea-specific genes
Yuzuru Sasamoto, Ryuhei Hayashi, Sung-Joon Park, Mihoko Saito-Adachi, Yutaka Suzuki, Satoshi Kawasaki, Andrew Quantock, Kenta Nakai, Motokazu Tsujikawa and Kohji Nishida
Investigative Ophthalmology & Visual Science (2016), 57(12):4906, ARVO Annual Meeting Abstract [IOVS] -
バイオインフォマティクス: 第9章 プロテオミクス
Andrzej Polanski (著), 後藤 修 (翻訳), 朴 聖俊 (第9章翻訳),丸善出版, 2012
-
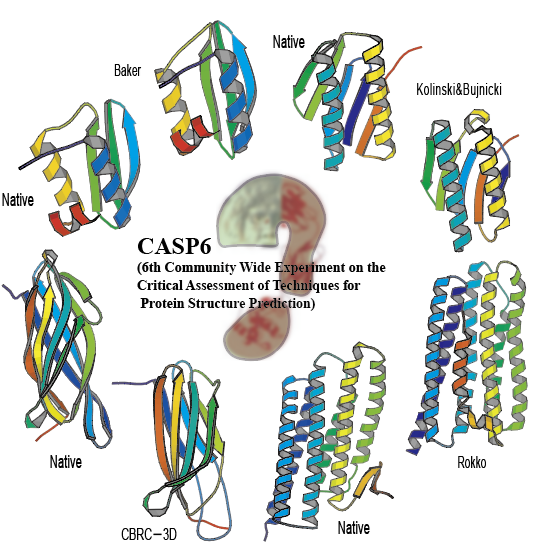
Template-free Prediction by Fragment Assembly with SimFold Energy Function at CASP7
S.J. Park, N. Hori, K. Okazaki and S. Takada
CASP7 Abstracts, 110-112, 2006 [PDF] [WEB] -
De novo Structure Prediction Server by Fragment Assembly with SimFold Energy Function
W. Jin, S.J. Park and S. Takada, CASP7 Abstracts, 112-113, 2006 [PDF]
-
タンパク質立体構造予測の現状と未来
朴 聖俊, 千見寺 浄慈, 広川 貴次, 富井 健太郎, 高田 彰二
人工知能学会誌, 20(4):479-485, 2005 [J-STAGE] -
タンパク質立体構造の2層比較
朴 聖俊, 山村 雅幸, 第48回 数理モデル化と問題解決(MPS)研究会, 2004 [CiNii]

|
|
|
|
|
||

|
|
|
|
|
||

|
|
|
|
|
||

|
|
|

|
|
|
|
|
||
|
| ||

|
|
|
|
|
||
|
|
||
 [cover image]
[cover image]
|
|
|
|
|
-
OmniTFT: Omni Target Forecasting for Vital Signs and Laboratory Result Trajectories in Multi Center ICU Data
Wanzhe Xu, Yutong Dai, Yitao Yang, Martin Loza, Weihang Zhang, Yang Cui, Xin Zeng, Sung Joon Park, Kenta Nakai
2025, arXiv:2511.19485 -
Extensive Computational Screening Identifies Marco (SR-A6) and Abca1/2 as Key Regulators of Intracellular Lipid Metabolism to Compensate Low Cholesterol Biosynthesis in M1 Macrophages Hysteresis
Yubo Zhang, Wenbo Yang, Yutaro Kumagai, Lopez Martin, Yitao Yang, Sung-Joon Park, Kenta Nakai
2024, doi: 10.20944/preprints202411.0082.v1 -
STAIG: Spatial Transcriptomics Analysis via Image-Aided Graph Contrastive Learning for Domain Exploration and Alignment-Free Integration
Yitao Yang, Yang Cui, Xin Zeng, Yubo Zhang, Martin Loza, Sung-Joon Park, Kenta Nakai
2023, bioRxiv 2023.12.18.572279, doi: https://doi.org/10.1101/2023.12.18.572279 -
HyGAnno: Hybrid graph neural network-based cell type annotation for single-cell ATAC sequencing data
Weihang Zhang, Yang Cui, Bowen Liu, Martin Loza Lopez, Sung-Joon Park, Kenta Nakai
2023, bioRxiv 2023.11.29.569114, doi: https://doi.org/10.1101/2023.11.29.569114 -
A computational approach for deciphering the interactions between proximal and distal regulators in B cell differentiation
Sung-Joon Park and Kenta Nakai
2023, bioRxiv 2023.11.02.565268, doi: https://doi.org/10.1101/2023.11.02.565268 -
Multi-omics computational analysis unveils the involvement of AP-1 and CTCF in hysteresis of chromatin states during macrophage polarization
Yubo Zhang, Wenbo Yang, Yutaro Kumagai, Martin Loza, Sung-Joon Park, Kenta Nakai
2023, bioRxiv 2023.09.27.559849, doi: https://doi.org/10.1101/2023.09.27.559849 -
RNA-sequencing reveals positional memory of multipotent mesenchymal stromal cells from oral and maxillofacial tissue transcriptomes
Satoru Onizuka, Yasuharu Yamazaki, Sung-Joon Park, Takayuki Sugimoto, Yumiko Sone, Sebastian Sjöqvist, Michihiko Usui, Akira Takeda, Kenta Nakai, Keisuke Nakashima, Takanori Iwata
2020, https://doi.org/10.21203/rs.2.24666/v2 -
Existence and possible roles of independent non-CpG methylation in mammalian brain
Jong-Hun Lee, Yutaka Saito, Sung-Joon Park, Kenta Nakai
2020, https://doi.org/10.21203/rs.2.22195/v1 -
A systematic NGS-based approach for contaminant detection and functional inference
Sung-Joon Park, Satoru Onizuka, Masahide Seki, Yutaka Suzuki, Takanori Iwata, Kenta Nakai
2019, bioRxiv 741934, doi: https://doi.org/10.1101/741934 -
RNA-Sequencing Reveals Unique Features of Multipotent Mesenchymal Stromal Cells from Oral and Maxillofacial Tissue Transcriptomes
Satoru Onizuka, Yasuharu Yamazaki, Sung-Joon Park, Takayuki Sugimoto, Yumiko Sone, Sebastian Sjöqvist, Michihiko Usui, Akira Takeda, Kenta Nakai, Keisuke Nakashima, Takanori Iwata
2019, http://dx.doi.org/10.2139/ssrn.3408026
RNAシーケンス解析による薬剤性顎骨壊死患者歯肉由来間葉系間質細胞の解析
西巻 和広, 貝淵 信之, 朴 聖俊, 橘川 眞一, 大和 雅之, 中井 謙太, 古賀 陽子, 岡本俊宏 第24回日本再生医療学会総会, P-05-07, Mar.20-22, 2025, 横浜
A graph-embedding approach to dissecting proximal and distal gene regulators
Sung-Joon Park 9th International Conference on Biomedical Engineering and Applications (ICBEA 2025), invited talk, Feb.27-Mar.02, 2025, Seoul, Korea
Identifying regulators of global chromatin accessibility using CRISPR-ATACsee-ATACseq
Sung-Joon Park and Yusuke Miyanari The 1st Asia & Pacific Bioinformatics Joint Conference (APBJC2024), 67/23-IV-2, Oct.22-24, 2024, Okinawa, Japan
STAIG: Spatial Transcriptomics Analysis via Image-Aided Graph Contrastive Learning for Domain Exploration and Alignment-Free Integration
Yitao Yang, Cui Yang, Xin Zeng, Yubo Zhang, Martin Loza, Sung-Joon Park and Kenta Nakai The 1st Asia & Pacific Bioinformatics Joint Conference (APBJC2024), 211/23-I-13, Oct.22-24, 2024, Okinawa, Japan
Integrative multimodal analysis identifying COPS5 as a novel biomarker for diffuse large B-cell lymphoma
Yutong Dai, Jingmei Li, Keita Yamamoto, Susumu Goyama, Martin Loza, Sung-Joon Park and Kenta Nakai The 1st Asia & Pacific Bioinformatics Joint Conference (APBJC2024), 24-XII-7, Oct.22-24, 2024, Okinawa, Japan
A computational approach for deciphering the interactions between proximal and distal gene regulators
Sung-Joon Park and Kenta Nakai 第76回日本細胞生物学会大会, 3A06, Jul.17-19, 2024, つくば国際会議場
STAIG: Spatial Transcriptomics Analysis via Image-Aided Graph Contrastive Learning for Domain Exploration and Alignment-Free Integration
Yitao Yang, Yang Cui, Xin Zeng, Yubo Zhang, Martin Loza Lopez, Sung-Joon Park, Kenta Nakai 第23回 東京大学生命科学シンポジウム (BIO UT 2024), P078, June. 21-22, 2024
mTOR/p53共標的治療は, 悪性度の高い急性骨髄単球性白血病に対して強い治療効果を示す
李 ショウ美, 杉本 絵美, 山本 圭太, 戴 彧童, 朴 聖俊, 中井 謙太, 北村 俊雄, 合山 進 日本薬学会第144年会, 30-503-pm02S, Mar.28-31, 2024, 横浜 学生優秀発表賞(口頭発表の部)
機械学習法によるびまん性B細胞リンパ腫の新規バイオマーカーCOPS5の同定
戴 彧童, 朴 聖俊, 李 婧美, ⼭本 圭太, 合⼭ 進, 中井 謙太
第46回日本分子生物学会年会, 1P-819, Dec. 6-8, 2023, 神戸
Exploring the role of hysteresis in macrophage lipid metabolism and repolarization memory
Yubo Zhang, Yutaro Kumagai, Sung-Joon Park and Kenta Nakai
The 22nd International Conference on Bioinformatics (InCoB 2023), Nov. 12-15, 2023, Brisbane, Australia
Identification of COPS5 as a novel biomarker of diffuse large B cell lymphoma by a machine-learning algorithm
Yutong Dai, Sung-Joon Park, Jingmei Li, Keita Yamamoto, Susumu Goyama and Kenta Nakai
The 22nd International Conference on Bioinformatics (InCoB 2023), Nov. 12-15, 2023, Brisbane, Australia
A graph-embedding approach dissecting enhancer-cofactor-promoter interactions
Sung-Joon Park and Kenta Nakai
The 22nd International Conference on Bioinformatics (InCoB 2023), Nov. 12-15, 2023, Brisbane, Australia
Computational modeling of the interactions between proximal and distal transcriptional regulators
Sung-Joon Park and Kenta Nakai
The 4th GENOME MODALITY Annual meeting, Sep. 21-23, 2023, Karolinska Institute, Sweden
A BERT-based prediction model of human protein sub-cellular localization
Wanzhe Xu, Sung-Joon Park, and Kenta Nakai
The 4th GENOME MODALITY Annual meeting, Sep. 21-23, 2023, Karolinska Institute, Sweden
Prediction of global splice-site selection considering long-range effects using Transformer
Yuna Miyachi, Sung-Joon Park, and Kenta Nakai
The 4th GENOME MODALITY Annual meeting, Sep. 21-23, 2023, Karolinska Institute, Sweden
長距離および短距離転写制御因子間相互作用の計算モデリング手法の開発
朴 聖俊, 中井 謙太
第12回生命医薬情報学連合大会 (IIBMP 2023), OS-3/P-46, Sep. 7-9, 2023, Kashiwa, Japan
BEST ORAL PRESENTATION AWARD
Meta co-expression analysis of genetic heterogeneity in GBM using single-cell RNA-seq data
Taisuke Hori, Sung-Joon Park and Kenta Nakai
生命医薬情報学連合大会 (IIBMP 2021), Sep. 27-29, 2021, Japan
BEST ORAL PRESENTATION AWARD
A Semi-supervised Deep Learning Approach for Predicting the Functional Effects of Genomic Non-coding Variations
Hao Jia, Sung-Joon Park and Kenta Nakai
The 19th International Conference on Bioinformatics (InCoB 2020), Oral, Nov. 25-29, 2020, VIRTUALLY
BEST PAPER PRESENTATION AWARD
Characterizing Promoter and Enhancer Sequences by a Deep Learning Method
Xin Zeng, Sung-Joon Park and Kenta Nakai
The 19th International Conference on Bioinformatics (InCoB 2020), Oral, Nov. 25-29, 2020, VIRTUALLY
Analyzing the impact of inter-chromosomal interactions on B cell development by linear regression modeling
Sung-Joon Park
生命医薬情報学連合大会 (IIBMP 2020), Oral O-6, Sep. 1-3, 2020, Japan
EXCELLENT ORAL PRESENTATION AWARD
Co-expression network analysis of E3 ligase genes in embryonic stem cells and cell fate specification
Darina Obukhova, Sung-Joon Park and Kenta Nakai
生命医薬情報学連合大会 (IIBMP 2020), Poster P-40, Sep. 1-3, 2020, Japan
歯根膜組織由来間葉系幹細胞シートによる歯周組織再生
鬼塚 理, 朴 聖俊, 貝淵 信之, 妻沼 有香, 安藤 智博, 中井 謙太, 岩田 隆紀
第19回日本再生医療学会総会, Mar. 12-14, 2020, PACIFICO Yokohama, Japan
Regression-based promoter modeling with local 3D chromatin structure
Sung-Joon Park
CHROMOSOME DYNAMICS (An international symposium on chromatin and chromosome stability), Dec. 8-10, 2019, Basel, Switzerland
脂肪由来幹細胞の運命決定と分化制御に関する時空間特異的ヒストン修飾のin-silico特定
山谷 恭代, 朴 聖俊, 中井 謙太
第42回 日本分子生物学会年会, 2P-0091, Dec. 3-6, 2019, 福岡, 日本
Genome-wide chromatin structural analysis reveals inter-chromosomal translocations in B-cell lymphoma
Luis A.E. Nagai, Sung-Joon Park, Kenta Nakai
The 42nd Annual Meeting of the Molecular Biology Society of Japan, 1P-0002, Dec. 3-6, 2019, Fukuoka, Japan
脂肪由来幹細胞の分化制御に働くエピジェネティック因子のin-silico同定
山谷 恭代, 朴 聖俊, 中井 謙太
IIBMP2019, O-1 (oral), Sep.9-11, 2019, Tokyo, Japan
Analyzing the 3D Chromatin Organization Coordinating with Gene Expression Regulation in B-cell Lymphoma
Luis Augusto Eijy Nagai, Sung-Joon Park and Kenta Nakai
IIBMP2019, O-3 (oral), Sep.9-11, 2019, Tokyo, Japan
OpenContami: A Web-based Application for Detecting Microbial Contaminants in Next-generation Sequencing
Sung-Joon Park, Satoru Onizuka, Masahide Seki, Yutaka Suzuki, Takanori Iwata and Kenta Nakai
IIBMP2019, O-10 (oral), Sep.9-11, 2019, Tokyo, Japan
Analyzing the 3D Chromatin Organization Coordinating with Gene Expression Regulation in B-cell Lymphoma
Luis Augusto Eijy Nagai, Sung-Joon Park and Kenta Nakai
IIBMP2019, P-1 (poster), Sep.9-11, 2019, Tokyo, Japan
脂肪由来幹細胞の分化制御における時期特異的ヒストン修飾のin-silico検出
山谷 恭代, 朴 聖俊, 中井 謙太
IIBMP2019, P-56 (poster), Sep.9-11, 2019, Tokyo, Japan
マウス胎仔期生殖細胞におけるクロマチンリプログラミング
山中 総一郎, 西原 秀典, 藤 英博, Luis Augusto Eijy Nagai, 橋本 浩介, 朴 聖俊, 渋谷 あおい, Ana Maria Suzuki, 田中 悠二郎, 中井 謙太, Piero Carninci, 佐々木 裕之, 塩見 春彦
第21回日本RNA学会年会, P-45, Jul.17-19, 2019, Tokyo, Japan
Host-unmapped NGS reads shape the prevalence of microbial contamination
Sung-Joon Park and Kenta Nakai
The Biology of Genomes, P188, May 7-11, 2019, Cold Spring Harbor Laboratory, NY, USA
Genome-wide chromatin structural analysis reveals inter-chromosomal translocations in B-cell lymphoma cell line
Luis AE Nagai, Sung-Joon Park and Kenta Nakai
The Biology of Genomes, P177, May 7-11, 2019, Cold Spring Harbor Laboratory, NY, USA
Profiling microbial contamination by exploiting host-unmapped NGS reads
Sung-Joon Park and Kenta Nakai
Revolutionizing Next-Generation Sequencing 2019 (RNGS19), P36, Mar. 25-26, 2019, Antwerp, Belgium
同種歯根膜由来間葉系幹細胞シートによる歯周組織の再建
岩田 隆紀, 鬼塚 理, 妻沼 有香, 朴 聖俊, 中井 謙太, 安藤 智博
第18回日本再生医療学会総会, SY-45-3, Mar. 21-23, 2019, Kobe, Japan
ゴノサイトでの新規DNAメチル化に付随して起こる多段階のクロマチンの緩み
山中 総一郎, 西原 秀典, 藤 英博, 永井 映司, 橋本 浩介, 朴 聖俊, 渋谷 あおい, Ana Maria Suzuki, 田中 悠二郎, 中井 謙太, Piero Carninci, 佐々木 裕之, 塩見 春彦
第41回日本分子生物学会年会, 3AW-09-1 (2P-0007), Nov. 28-30, 2018, Yokohama, Japan
Applying Joint Non-Negative Matrix Factorization to Functional Analysis of Microbial Infection
Sung-Joon Park and Kenta Nakai
The 17th International Conference On BioInformatics (InCoB-2018), P-55, Sep. 26-28, 2018, New Delhi, India
Analyzing the 3D Chromatin Organization Coordinating with Gene Expression Regulation in B-cell Lymphoma
Luis A.E. Nagai, Sung-Joon Park and Kenta Nakai
The 17th International Conference On BioInformatics (InCoB-2018), Sep. 26-28, 2018, New Delhi, India
BEST ORAL PRESENTATION AWARD
Three-Dimensional Chromatin Structures Coordinate with Gene Expression Levels and Reveals Inter-chromosomal Interactions in B-Cell Lymphoma
Luis A.E. Nagai, Sung-Joon Park and Kenta Nakai
生命医薬情報学連合大会 (IIBMP2018), P-31 (oral presentation), 2018年9月19日~21日, 鶴岡、日本
OpenLooper: A web-based platform for managing multi-layered NGS data
Sung-Joon Park and Kenta Nakai
生命医薬情報学連合大会 (IIBMP2018), P-32, 2018年9月19日~21日, 鶴岡、日本
脂肪由来幹細胞(Adipose-derived stem cell)の分化に関与するエピジェネティック因子の探索
山谷 恭代, 朴 聖俊, 中井 謙太
生命医薬情報学連合大会 (IIBMP2018), P-37 (oral presentation), 2018年9月19日~21日, 鶴岡、日本
DNAヒドロキシメチル化とヒストンH4K8アセチル化によるヒト多能性幹細胞遺伝子発現変動
石川 靖久, 朴 聖俊, 中井 謙太
生命医薬情報学連合大会 (IIBMP2018), P-73, 2018年9月19日~21日, 鶴岡、日本
The establishment of safety and efficacy evaluation for allogeneic periodontal ligament derived multipotent mesenchymal stromal cell sheet with next-generation sequencer
Takanori Iwata, Satoru Onizuka, Sung-Joon Park, Yuka Tsumanuma, Kenta Nakai, Yuichi Izumi, Tomohiro Ando
5th TERMIS World Congress, 36-SY-2, Sep. 4-7, 2018, Kyoto, Japan
Detecting Microbial Contamination from NGS Data and Analyzing Its Functional Impact
Sung-Joon Park and Kenta Naka
BIO UT, P70, June 9, 2018, Tokyo, Japan
Profiling of Cell Microbial Contamination by NGS data
Sung-Joon Park 配列解析シンポジウム (36 years since the Smith-Waterman-Gotoh algorithm), 2018年3月28日,産業技術総合研究所 臨海副都心センター別館
同種歯根膜細胞シートの安全性・有効性評価指標の確立と歯周組織の再建
岩田 隆紀, 鬼塚 理, 朴 聖俊, 妻沼 有香, 和泉 雄一, 中井 謙太, 安藤 智博
第17回 日本再生医療学会総会, 2018年3月21-23日, 横浜, 日本
Non-CpGメチル化パターンによる脳の老化関連遺伝子の分類
李 鍾勳, 朴 聖俊, 中井 謙太
2017年度生命科学系学会合同年次大会, 1P-0670, 2017年12月6日, 神戸, 日本
生殖細胞においてトランスポゾンに富んだ染色体領域が一過的に緩む
山中 総一郎, Sung-Joon Park, 渋谷 あおい, Piero Carninci, 中井 謙太, 塩見 春彦
2017年度生命科学系学会合同年次大会, 3P-0559 (3PT18-13), 2017年12月8日, 神戸, 日本
Development of a Pipeline for Contamination Profiling of Cells with Next Generation Sequencing Data
Sung-Joon Park, Satoru Onizuka, Takanori Iwata, Kenta Nakai
The 28th International Conference on Genome Informatics Workshop (GIW)/BIOINFO 2017, P35, Oct.31-Nov.3, 2017, Seoul, Korea
BEST POSTER AWARD
Whole transcriptome analysis of MSCs derived from different types of tissue reveals unique profiles
Satoru Onizuka, Yasuharu Yamazaki, Takayuki Sugimoto, Yumiko Sone, Akira Takeda, Sung-Joon Park, Kenta Nakai, Takanori Iwata, Masayuki Yamato, Teruo Okano
The American Society for Bone and Mineral Research (ASBMR) 2017 Annual Meeting, Sep.8-11, 2017, Denver, Colorado, USA
精子エピジェネティックマップ:その本質とリプログラミング様式の解明に向けて
岡田 由紀, 山口 幸佑, 牧野 吉倫, 朴 聖俊, 加藤 由起, 中井 謙太, 白髭 克彦
第44回日本毒性学会学術年会, 2017年7月10日-12日, S6-3, 横浜, 日本
腸骨・顎骨組織由来間葉系幹細胞に対する網羅的遺伝子発現解析による検討
鬼塚 理, 山崎安晴, 杉本孝之, 曽根由美子, 武田 啓, 朴 聖俊, 中井謙太, 岩田隆紀, 大和雅之, 岡野光夫
第16回日本再生医療学会総会, 2017年3月7日, 仙台, 日本
歯根膜細胞シートを用いた歯周組織の再生と展望
岩田隆紀, 鬼塚 理, 伊豆原 るな, 鷲尾 薫, 妻沼有香, Park Sung-Joon, 中井謙太, 和泉雄一, 安藤智博
第16回日本再生医療学会総会, 2017年3月8日, 仙台, 日本
An Empirical Model for Visualizing Clonal Expansion of HTLV-1-Infected Cells in Adult T-Cell Leukemia Derived from High-Throughput Sequencing
Amir Farmanbar, Sanaz Firouzi, Sung-Joon Park, Kaoru Uchimaru, Toshiki Watanabe, Kenta Nakai
18th International Conference on Human Retrovirology HTLV and Related Viruses, March 7-10 2017, Tokyo, Japan
BEST POSTER AWARD
Comprehensive Clonality Analysis of HTLV-1 Infected Cells Integrating Cell Surface Markers of ATL Progression, Genome Wide Profiling of Provirus Integration Sites and Mutation Patterns
Sanaz Firouzi, Amir Farmanbar, Sereewattanawoot Sarun (Ball), Seiichiro Kobayashi,Kazumi Nakano, Sung-Joon Park, Kenta Nakai, Toshiki Watanabe, Yutaka Suzuki, Kaoru Uchimaru
18th International Conference on Human Retrovirology HTLV and Related Viruses, March 7-10 2017, Tokyo, Japan
Assessing the Impact of Contamination on RNA-seq Transcriptome Profiles
Sung-Joon Park, Josep Basha Gutierrez, Myungjin Moon, Kenta Nakai
The 15th International Conference On BioInformatics (InCOB 2016), Sep.21-23, Biopolis, Singapore
BEST POSTER AWARD (GOLD)
多施設培養幹細胞のトランスクリプトーム比較解析
朴 聖俊, 中井 謙太
BMB2015 (第38回日本分子生物学会年会、第88回日本生化学会大会 合同大会), 2W12-p-4, Dec.1-4, 2015, Kobe, Japan
次世代シークエンサーによる再生医療のためのヒト間葉系幹細胞の品質管理
岩田 隆紀, 朴 聖俊, 各務 秀明, 中井 謙太, 大和 雅之
BMB2015 (第38回日本分子生物学会年会、第88回日本生化学会大会 合同大会), 2W12-p-2, Dec.1-4, 2015, Kobe, Japan
ゲノム編集した患者由来疾患iPS細胞の全ゲノム解析によって明らかになったこと
曽根 岳史, 朴 聖俊, 田中 泰圭, 太田 悦朗, 日暮 憲道, 中井 謙太, 廣瀬 伸一, 岡野 栄之
BMB2015 (第38回日本分子生物学会年会、第88回日本生化学会大会 合同大会), 2W12-p-3, Dec.1-4, 2015, Kobe, Japan
Integrin αvβ3 による双方向的な造血幹細胞活性の制御
梅本 晃正, 松崎 優, 大和 雅之, 古澤 純一, 善本 隆之, 朴 聖俊, 中井 謙太, 須田 年生
BMB2015 (第38回日本分子生物学会年会、第88回日本生化学会大会 合同大会), 2W12-p-5, Dec.1-4, 2015, Kobe, Japan
OpenTein: a database of whole-slide digital images of stem cell teratomas
Sung-Joon Park, Yusuke Komiyama, Hirofumi Suemori, Akihiro Umezawa, Kenta Nakai
GIW/InCoB 2015, P166, Sept.9-11, 2015, Odaiba, Tokyo
Analysis of differential regulatory network dynamics in hematopoietic stem and progenitor cells
Sung-Joon Park, Terumasa Umemoto, Mihoko Saito-Adachi, Masayuki Yamato, Kenta Nakai
Annual Conference of Japanese Society for Bioinformatics, P48, 2013
Inferring gene regulatory network from RNA-seq data of mouse hematopoietic stem and progenitor cells
Sung-Joon Park, Terumasa Umemoto, Masayuki Yamato, Kenta Nakai
The 8th International Symposium of the Institute Network, P1, 2013
タウリンによるTXNIP発現制御に関する研究
権藤 祐輔, 薩 秀夫, 朴 聖俊, 若林 俊一, 中井 謙太, 清水 誠
日本農芸化学会2013年度大会, 3月24-28日, 仙台, 2013
Massive-scale RNA-Seq analysis of mouse early embryo development
Sung-Joon Park, Makiko Komata, Fukashi Inoue, Kaori Yamada, Kenta Nakai, Miho Ohsugi, Katsuhiko Shirahige,
The 22nd International Conference on Genomic Informatics (GIW 2011), Posters and Software Demonstrations, P-017, Dec 5-7, Busan, Korea, 2011
OUTSTANDING POSTER AWARD
Transcriptome analysis of mouse early embryo development by RNA sequencing
Sung-Joon Park, Makiko Komata, Fukashi Inoue, Kaori Yamada, Kenta Nakai, Miho Ohsugi, Katsuhiko Shirahige
Annual Conference of Japanese Society for Bioinformatics, Selected oral presentation (JSBi-46), 2011
Predicting Promoter Activities of Primary Human DNA Sequences
Takuma Irie, Sung-Joon Park, Riu Yamashita, Masahide Seki, Tetsushi Yada, Sumio Sugano, Yutaka Suzuki
The 34th Annual Meeting of the Molecular Biology Society of Japan, 3P-0196, 2011
Predicting Promoter Activities of Primary Human DNA Sequences
Takuma Irie, Sung-Joon Park, Riu Yamashita, Masahide Seki, Tetsushi Yada, Sumio Sugano, Kenta Nakai, Yutaka Suzuki,
Biochemistry and Molecular Biology (BMB2010), 3P-0694, 2010
真核生物の遺伝子転写制御機構のモデル化と解析
朴 聖俊
九州工業大学情報工学部生命情報工学科講演会 (Jan.28), 2010
大規模遺伝子プロモータ活性におけるシス制御因子の重要度推定
朴 聖俊
第41回 人工知能学会 分子生物情報研究会, 2009
Comprehensive Modeling for the Combinatorial Regulation of Transcription Factors
Sung-Joon Park, Natsuhiro Ichinose and Tetsushi Yada
Annual Conference of Japanese Society for Bioinformatics, Selected oral presentation (P006/T08), 2008
Template-free Predictionの現状と今後
朴 聖俊
第7回GSICシンポジウム (34回分子生物情報研究会), 2006
粗視化モデルによるタンパク質の局所構造変化とトポロジー形成との相関解析
朴 聖俊
第16回 生命情報科学特別講義(CBRC), 2005.9.26
アミノ酸配列情報からの蛋白質立体構造予測へ向けて:配列プロファイル比較に基づく部分立体構造ライブラリの構築
朴 聖俊, 高田 彰二
第32回知能システムシンポジウム, 289-294, 2005
タンパク質立体構造予測技術の現状と未来:Rokko, Rokky 神戸大学理学部チーム
高田 章二, 朴 聖俊
IPAB公開セミナー, 2005
フラグメントアセンブリ法によるタンパク質立体構造予測のための戦略的フラグメント探索
朴 聖俊, 千見寺 浄慈, 藤墳 佳見, 高田 彰二
日本生物物理学会第42回年会, 2P042, 2004
タンパク質立体構造予測サーバーROKKYの開発
朴 聖俊, 金 文珍,古田 忠臣, 高田 彰二
バイオグリッド研究会, 2004
非同期並列GAによるタンパク質の立体構造アラインメント
朴 聖俊,山村 雅幸
分子プログラミング夏季セミナー, 2003
ローカルとグローバル立体構造比較から眺めるタンパク質進化
朴 聖俊,山村 雅幸
第24回 SIG-MBI研究会, 2003
Protein Structure-based Alignment - Optimized Meaningful Fragment Posture by Real-coded Genetic Algorithm -
Sung-Joon Park and Masayuki Yamamura
SICE Sympossium on Systems and Information 2002, 269-272, 2002
Comparative Sensitivity of Protein Structure-based Alignment Algorithms: An Essential Approach to Identifying Positional Conservation
Sung-Joon Park and Masayuki Yamamura
SICE The 29th Intelligent System Symposium, 65-68, 2002
二つのタンパク質における構造的類似度を発見するための遺伝的アルゴリズムの適用
朴 聖俊,山村 雅幸
第10回 SIG-MBI研究会, 2000
遺伝的立体構造アラインメント:マルチプル立体構造アラインメントへ向けて
朴 聖俊,山村 雅幸
SICE第27回知能システムシンポジウム, 123-126, 2000
GAによる立体構造アラインメント
朴 聖俊,山村 雅幸
SICE第12回自律分散システム, 323-328, 2000